This HTML version of
You might prefer to read the PDF version, or you can buy a hard copy from Amazon.
Chapter 10 Approximate Bayesian Computation
10.1 The Variability Hypothesis
I have a soft spot for crank science. Recently I visited Norumbega Tower, which is an enduring monument to the crackpot theories of Eben Norton Horsford, inventor of double-acting baking powder and fake history. But that’s not what this chapter is about.
This chapter is about the Variability Hypothesis, which
"originated in the early nineteenth century with Johann Meckel, who argued that males have a greater range of ability than females, especially in intelligence. In other words, he believed that most geniuses and most mentally retarded people are men. Because he considered males to be the ’superior animal,’ Meckel concluded that females’ lack of variation was a sign of inferiority."
I particularly like that last part, because I suspect that if it turns out that women are actually more variable, Meckel would take that as a sign of inferiority, too. Anyway, you will not be surprised to hear that the evidence for the Variability Hypothesis is weak.
Nevertheless, it came up in my class recently when we looked at data from the CDC’s Behavioral Risk Factor Surveillance System (BRFSS), specifically the self-reported heights of adult American men and women. The dataset includes responses from 154407 men and 254722 women. Here’s what we found:
- The average height for men is 178 cm; the average height for women is 163 cm. So men are taller, on average. No surprise there.
- For men the standard deviation is 7.7 cm; for women it is 7.3 cm. So in absolute terms, men’s heights are more variable.
- But to compare variability between groups, it is more meaningful to use the coefficient of variation (CV), which is the standard deviation divided by the mean. It is a dimensionless measure of variability relative to scale. For men CV is 0.0433; for women it is 0.0444.
That’s very close, so we could conclude that this dataset provides weak evidence against the Variability Hypothesis. But we can use Bayesian methods to make that conclusion more precise. And answering this question gives me a chance to demonstrate some techniques for working with large datasets.
I will proceed in a few steps:
- We’ll start with the simplest implementation, but it only works for datasets smaller than 1000 values.
- By computing probabilities under a log transform, we can scale up to the full size of the dataset, but the computation gets slow.
- Finally, we speed things up substantially with Approximate Bayesian Computation, also known as ABC.
You can download the code in this chapter from http://thinkbayes.com/variability.py. For more information see Section 0.3.
10.2 Mean and standard deviation
In Chapter 9 we estimated two parameters simultaneously using a joint distribution. In this chapter we use the same method to estimate the parameters of a Gaussian distribution: the mean, mu, and the standard deviation, sigma.
For this problem, I define a Suite called Height that represents a map from each mu, sigma pair to its probability:
class Height(thinkbayes.Suite, thinkbayes.Joint):
def __init__(self, mus, sigmas):
pairs = [(mu, sigma)
for mu in mus
for sigma in sigmas]
thinkbayes.Suite.__init__(self, pairs)
mus is a sequence of possible values for mu; sigmas is a sequence of values for sigma. The prior distribution is uniform over all mu, sigma pairs.
The likelihood function is easy. Given hypothetical values of mu and sigma, we compute the likelihood of a particular value, x. That’s what EvalGaussianPdf does, so all we have to do is use it:
# class Height
def Likelihood(self, data, hypo):
x = data
mu, sigma = hypo
like = thinkbayes.EvalGaussianPdf(x, mu, sigma)
return like
If you have studied statistics from a mathematical perspective, you know that when you evaluate a PDF, you get a probability density. In order to get a probability, you have to integrate probability densities over some range.
But for our purposes, we don’t need a probability; we just need something proportional to the probability we want. A probability density does that job nicely.
The hardest part of this problem turns out to be choosing appropriate ranges for mus and sigmas. If the range is too small, we omit some possibilities with non-negligible probability and get the wrong answer. If the range is too big, we get the right answer, but waste computational power.
So this is an opportunity to use classical estimation to make Bayesian techniques more efficient. Specifically, we can use classical estimators to find a likely location for mu and sigma, and use the standard errors of those estimates to choose a likely spread.
If the true parameters of the distribution are µ and σ, and we take a sample of n values, an estimator of µ is the sample mean, m.
And an estimator of σ is the sample standard variance, s.
The standard error of the estimated µ is s / √n and the standard error of the estimated σ is s / √2 (n−1).
Here’s the code to compute all that:
def FindPriorRanges(xs, num_points, num_stderrs=3.0):
# compute m and s
n = len(xs)
m = numpy.mean(xs)
s = numpy.std(xs)
# compute ranges for m and s
stderr_m = s / math.sqrt(n)
mus = MakeRange(m, stderr_m, num_stderrs)
stderr_s = s / math.sqrt(2 * (n-1))
sigmas = MakeRange(s, stderr_s, num_stderrs)
return mus, sigmas
xs is the dataset. num_points is the desired number of
values in the range. num_stderrs is the width of the range on
each side of the estimate, in number of standard errors.
The return value is a pair of sequences, mus and sigmas.
def MakeRange(estimate, stderr, num_stderrs):
spread = stderr * num_stderrs
array = numpy.linspace(estimate-spread,
estimate+spread,
num_points)
return array
numpy.linspace makes an array of equally spaced elements between estimate-spread and estimate+spread, including both.
10.3 Update
Finally here’s the code to make and update the suite:
mus, sigmas = FindPriorRanges(xs, num_points)
suite = Height(mus, sigmas)
suite.UpdateSet(xs)
print suite.MaximumLikelihood()
This process might seem bogus, because we use the data to choose the range of the prior distribution, and then use the data again to do the update. In general, using the same data twice is, in fact, bogus.
But in this case it is ok. Really. We use the data to choose the
range for the prior, but only to avoid computing a lot of
probabilities that would have been very small anyway. With
num_stderrs=4, the range is big enough to cover all values with
non-negligible likelihood. After that, making it bigger has no effect
on the results.
In effect, the prior is uniform over all values of mu and sigma, but for computational efficiency we ignore all the values that don’t matter.
10.4 The posterior distribution of CV
Once we have the posterior joint distribution of mu and sigma, we can compute the distribution of CV for men and women, and then the probability that one exceeds the other.
To compute the distribution of CV, we enumerate pairs of mu and sigma:
def CoefVariation(suite):
pmf = thinkbayes.Pmf()
for (mu, sigma), p in suite.Items():
pmf.Incr(sigma/mu, p)
return pmf
Then we use thinkbayes.PmfProbGreater to compute the
probability that men are more variable.
The analysis itself is simple, but there are two more issues we have to deal with:
- As the size of the dataset increases, we run into a series of computational problems due to the limitations of floating-point arithmetic.
- The dataset contains a number of extreme values that are almost certainly errors. We will need to make the estimation process robust in the presence of these outliers.
The following sections explain these problems and their solutions.
10.5 Underflow
If we select the first 100 values from the BRFSS dataset and run the analysis I just described, it runs without errors and we get posterior distributions that look reasonable.
If we select the first 1000 values and run the program again, we get
an error in Pmf.Normalize:
ValueError: total probability is zero.
The problem is that we are using probability densities to compute likelihoods, and densities from continuous distributions tend to be small. And if you take 1000 small values and multiply them together, the result is very small. In this case it is so small it can’t be represented by a floating-point number, so it gets rounded down to zero, which is called underflow. And if all probabilities in the distribution are 0, it’s not a distribution any more.
A possible solution is to renormalize the Pmf after each update, or after each batch of 100. That would work, but it would be slow.
A better alternative is to compute likelihoods under a log transform. That way, instead of multiplying small values, we can add up log likelihoods. Pmf provides methods Log, LogUpdateSet and Exp to make this process easy.
Log computes the log of the probabilities in a Pmf:
# class Pmf
def Log(self):
m = self.MaxLike()
for x, p in self.d.iteritems():
if p:
self.Set(x, math.log(p/m))
else:
self.Remove(x)
Before applying the log transform Log uses MaxLike to find m, the highest probability in the Pmf. It divide all probabilities by m, so the highest probability gets normalized to 1, which yields a log of 0. The other log probabilities are all negative. If there are any values in the Pmf with probability 0, they are removed.
While the Pmf is under a log transform, we can’t use Update, UpdateSet, or Normalize. The result would be nonsensical; if you try, Pmf raises an exception. Instead, we have to use LogUpdate and LogUpdateSet.
Here’s the implementation of LogUpdateSet:
# class Suite
def LogUpdateSet(self, dataset):
for data in dataset:
self.LogUpdate(data)
LogUpdateSet loops through the data and calls LogUpdate:
# class Suite
def LogUpdate(self, data):
for hypo in self.Values():
like = self.LogLikelihood(data, hypo)
self.Incr(hypo, like)
LogUpdate is just like Update except that it calls LogLikelihood instead of Likelihood, and Incr instead of Mult.
Using log-likelihoods avoids the problem with underflow, but while the Pmf is under the log transform, there’s not much we can do with it. We have to use Exp to invert the transform:
# class Pmf
def Exp(self):
m = self.MaxLike()
for x, p in self.d.iteritems():
self.Set(x, math.exp(p-m))
If the log-likelihoods are large negative numbers, the resulting likelihoods might underflow. So Exp finds the maximum log-likelihood, m, and shifts all the likelihoods up by m. The resulting distribution has a maximum likelihood of 1. This process inverts the log transform with minimal loss of precision.
10.6 Log-likelihood
Now all we need is LogLikelihood.
# class Height
def LogLikelihood(self, data, hypo):
x = data
mu, sigma = hypo
loglike = scipy.stats.norm.logpdf(x, mu, sigma)
return loglike
norm.logpdf computes the log-likelihood of the Gaussian PDF.
Here’s what the whole update process looks like:
suite.Log()
suite.LogUpdateSet(xs)
suite.Exp()
suite.Normalize()
To review, Log puts the suite under a log transform. LogUpdateSet calls LogUpdate, which calls LogLikelihood. LogUpdate uses Pmf.Incr, because adding a log-likelihood is the same as multiplying by a likelihood.
After the update, the log-likelihoods are large negative numbers, so Exp shifts them up before inverting the transform, which is how we avoid underflow.
Once the suite is transformed back, the probabilities are “linear” again, which means “not logarithmic”, so we can use Normalize again.
Using this algorithm, we can process the entire dataset without underflow, but it is still slow. On my computer it might take an hour. We can do better.
10.7 A little optimization
This section uses math and computational optimization to speed things up by a factor of 100. But the following section presents an algorithm that is even faster. So if you want to get right to the good stuff, feel free to skip this section.
Suite.LogUpdateSet calls LogUpdate once for each data point. We can speed it up by computing the log-likelihood of the entire dataset at once.
We’ll start with the Gaussian PDF:
| exp | ⎡ ⎢ ⎢ ⎢ ⎢ ⎢ ⎣ | − |
| ⎛ ⎜ ⎜ ⎝ |
| ⎞ ⎟ ⎟ ⎠ |
| ⎤ ⎥ ⎥ ⎥ ⎥ ⎥ ⎦ |
and compute the log (dropping the constant term):
| −logσ − |
| ⎛ ⎜ ⎜ ⎝ |
| ⎞ ⎟ ⎟ ⎠ |
|
Given a sequence of values, xi, the total log-likelihood is
| −logσ − |
| ⎛ ⎜ ⎜ ⎝ |
| ⎞ ⎟ ⎟ ⎠ |
|
Pulling out the terms that don’t depend on i, we get
| −n logσ − |
|
| (xi − µ)2 |
which we can translate into Python:
# class Height
def LogUpdateSetFast(self, data):
xs = tuple(data)
n = len(xs)
for hypo in self.Values():
mu, sigma = hypo
total = Summation(xs, mu)
loglike = -n * math.log(sigma) - total / 2 / sigma**2
self.Incr(hypo, loglike)
By itself, this would be a small improvement, but it creates an opportunity for a bigger one. Notice that the summation only depends on mu, not sigma, so we only have to compute it once for each value of mu.
To avoid recomputing, I factor out a function that computes the summation, and memoize it so it stores previously computed results in a dictionary (see http://en.wikipedia.org/wiki/Memoization):
def Summation(xs, mu, cache={}):
try:
return cache[xs, mu]
except KeyError:
ds = [(x-mu)**2 for x in xs]
total = sum(ds)
cache[xs, mu] = total
return total
cache stores previously computed sums. The try statement returns a result from the cache if possible; otherwise it computes the summation, then caches and returns the result.
The only catch is that we can’t use a list as a key in the cache, because it is not a hashable type. That’s why LogUpdateSetFast converts the dataset to a tuple.
This optimization speeds up the computation by about a factor of 100, processing the entire dataset (154 407 men and 254 722 women) in less than a minute on my not-very-fast computer.
10.8 ABC
But maybe you don’t have that kind of time. In that case, Approximate Bayesian Computation (ABC) might be the way to go. The motivation behind ABC is that the likelihood of any particular dataset is:
- Very small, especially for large datasets, which is why we had to use the log transform,
- Expensive to compute, which is why we had to do so much optimization, and
- Not really what we want anyway.
We don’t really care about the likelihood of seeing the exact dataset we saw. Especially for continuous variables, we care about the likelihood of seeing any dataset like the one we saw.
For example, in the Euro problem, we don’t care about the order of the coin flips, only the total number of heads and tails. And in the locomotive problem, we don’t care about which particular trains were seen, only the number of trains and the maximum of the serial numbers.
Similarly, in the BRFSS sample, we don’t really want to know the probability of seeing one particular set of values (especially since there are hundreds of thousands of them). It is more relevant to ask, “If we sample 100,000 people from a population with hypothetical values of µ and σ, what would be the chance of collecting a sample with the observed mean and variance?”
For samples from a Gaussian distribution, we can answer this question efficiently because we can find the distribution of the sample statistics analytically. In fact, we already did it when we computed the range of the prior.
If you draw n values from a Gaussian distribution with parameters µ and σ, and compute the sample mean, m, the distribution of m is Gaussian with parameters µ and σ / √n.
Similarly, the distribution of the sample standard deviation, s, is Gaussian with parameters σ and σ / √2 (n−1).
We can use these sample distributions to compute the likelihood of the
sample statistics, m and s, given hypothetical values
for µ and σ. Here’s a new version of LogUpdateSet
that does it:
def LogUpdateSetABC(self, data):
xs = data
n = len(xs)
# compute sample statistics
m = numpy.mean(xs)
s = numpy.std(xs)
for hypo in sorted(self.Values()):
mu, sigma = hypo
# compute log likelihood of m, given hypo
stderr_m = sigma / math.sqrt(n)
loglike = EvalGaussianLogPdf(m, mu, stderr_m)
#compute log likelihood of s, given hypo
stderr_s = sigma / math.sqrt(2 * (n-1))
loglike += EvalGaussianLogPdf(s, sigma, stderr_s)
self.Incr(hypo, loglike)
On my computer this function processes the entire dataset in about a second, and the result agrees with the exact result with about 5 digits of precision.
10.9 Robust estimation
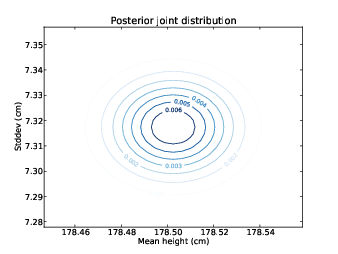
Figure 10.1: Contour plot of the posterior joint distribution of mean and standard deviation of height for men in the U.S.
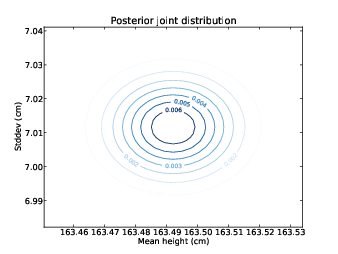
Figure 10.2: Contour plot of the posterior joint distribution of mean and standard deviation of height for women in the U.S.
We are almost ready to look at results, but we have one more problem to deal with. There are a number of outliers in this dataset that are almost certainly errors. For example, there are three adults with reported height of 61 cm, which would place them among the shortest living adults in the world. At the other end, there are four women with reported height 229 cm, just short of the tallest women in the world.
It is not impossible that these values are correct, but it is unlikely, which makes it hard to know how to deal with them. And we have to get it right, because these extreme values have a disproportionate effect on the estimated variability.
Because ABC is based on summary statistics, rather than the entire dataset, we can make it more robust by choosing summary statistics that are robust in the presence of outliers. For example, rather than use the sample mean and standard deviation, we could use the median and inter-quartile range (IQR), which is the difference between the 25th and 75th percentiles.
More generally, we could compute an inter-percentile range (IPR) that spans any given fraction of the distribution, p:
def MedianIPR(xs, p):
cdf = thinkbayes.MakeCdfFromList(xs)
median = cdf.Percentile(50)
alpha = (1-p) / 2
ipr = cdf.Value(1-alpha) - cdf.Value(alpha)
return median, ipr
xs is a sequence of values. p is the desired range; for example, p=0.5 yields the inter-quartile range.
MedianIPR works by computing the CDF of xs, then extracting the median and the difference between two percentiles.
We can convert from ipr to an estimate of sigma using the Gaussian CDF to compute the fraction of the distribution covered by a given number of standard deviations. For example, it is a well-known rule of thumb that 68% of a Gaussian distribution falls within one standard deviation of the mean, which leaves 16% in each tail. If we compute the range between the 16th and 84th percentiles, we expect the result to be 2 * sigma. So we can estimate sigma by computing the 68% IPR and dividing by 2.
More generally we could use any number of sigmas. MedianS performs the more general version of this computation:
def MedianS(xs, num_sigmas):
half_p = thinkbayes.StandardGaussianCdf(num_sigmas) - 0.5
median, ipr = MedianIPR(xs, half_p * 2)
s = ipr / 2 / num_sigmas
return median, s
Again, xs is the sequence of values; num_sigmas is the
number of standard deviations the results should be based on. The
result is median, which estimates µ, and s, which
estimates σ.
Finally, in LogUpdateSetABC we can replace the sample mean and standard deviation with median and s. And that pretty much does it.
It might seem odd that we are using observed percentiles to estimate µ and σ, but it is an example of the flexibility of the Bayesian approach. In effect we are asking, “Given hypothetical values for µ and σ, and a sampling process that has some chance of introducing errors, what is the likelihood of generating a given set of sample statistics?”
We are free to choose any sample statistics we like, up to a point: µ and σ determine the location and spread of a distribution, so we need to choose statistics that capture those characteristics. For example, if we chose the 49th and 51st percentiles, we would get very little information about spread, so it would leave the estimate of σ relatively unconstrained by the data. All values of sigma would have nearly the same likelihood of producing the observed values, so the posterior distribution of sigma would look a lot like the prior.
10.10 Who is more variable?
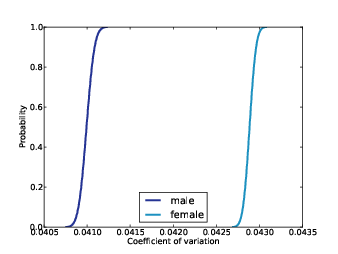
Figure 10.3: Posterior distributions of CV for men and women, based on robust estimators.
Finally we are ready to answer the question we started with: is the coefficient of variation greater for men than for women?
Using ABC based on the median and IPR with num_sigmas=1, I
computed posterior joint distributions for mu and sigma. Figures 10.1 and 10.2
show the results as a contour plot with mu on the x-axis, sigma on the y-axis, and probability on the z-axis.
For each joint distribution, I computed the posterior distribution of CV. Figure 10.3 shows these distributions for men and women. The mean for men is 0.0410; for women it is 0.0429. Since there is no overlap between the distributions, we conclude with near certainty that women are more variable in height than men.
So is that the end of the Variability Hypothesis? Sadly, no. It turns
out that this
result depends on the choice of the
inter-percentile range. With num_sigmas=1, we conclude that
women are more variable, but with num_sigmas=2 we conclude
with equal confidence that men are more variable.
The reason for the difference is that there are more men of short stature, and their distance from the mean is greater.
So our evaluation of the Variability Hypothesis depends on the
interpretation of “variability.” With num_sigmas=1 we
focus on people near the mean. As we increase
num_sigmas, we give more weight to the extremes.
To decide which emphasis is appropriate, we would need a more precise statement of the hypothesis. As it is, the Variability Hypothesis may be too vague to evaluate.
Nevertheless, it helped me demonstrate several new ideas and, I hope you agree, it makes an interesting example.
10.11 Discussion
There are two ways you might think of ABC. One interpretation is that it is, as the name suggests, an approximation that is faster to compute than the exact value.
But remember that Bayesian analysis is always based on modeling decisions, which implies that there is no “exact” solution. For any interesting physical system there are many possible models, and each model yields different results. To interpret the results, we have to evaluate the models.
So another interpretation of ABC is that it represents an alternative model of the likelihood. When we compute p(D|H), we are asking “What is the likelihood of the data under a given hypothesis?”
For large datasets, the likelihood of the data is very small, which is a hint that we might not be asking the right question. What we really want to know is the likelihood of any outcome like the data, where the definition of “like” is yet another modeling decision.
The underlying idea of ABC is that two datasets are alike if they yield the same summary statistics. But in some cases, like the example in this chapter, it is not obvious which summary statistics to choose.
You can download the code in this chapter from http://thinkbayes.com/variability.py. For more information see Section 0.3.
10.12 Exercises
An “effect size” is a statistic intended to measure the difference between two groups (see http://en.wikipedia.org/wiki/Effect_size).
For example, we could use data from the BRFSS to estimate the difference in height between men and women. By sampling values from the posterior distributions of µ and σ, we could generate the posterior distribution of this difference.
But it might be better to use a dimensionless measure of effect size, rather than a difference measured in cm. One option is to use divide through by the standard deviation (similar to what we did with the coefficient of variation).
If the parameters for Group 1 are (µ1, σ1), and the parameters for Group 2 are (µ2, σ2), the dimensionless effect size is
|
Write a function that takes joint distributions of mu and sigma for two groups and returns the posterior distribution of effect size.
Hint: if enumerating all pairs from the two distributions takes too long, consider random sampling.